Research Interest
Our lab is interested in building technologies that solve critical research bottlenecks and answer fundamental biological questions. Below you'll see a few of our ongoing projects and their applications.
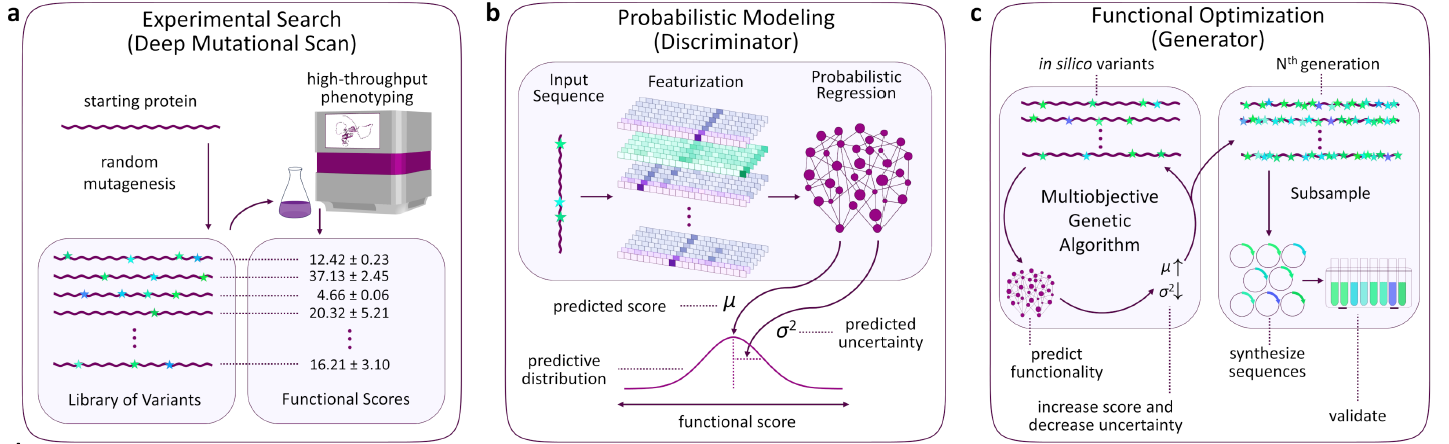
Combining high-throughput protein characterization with deep probabilistic modeling.
Designing proteins with desired functionality is a fundamental challenge of protein engineering. Previous efforts to rationally engineer protein variants often require a holistic understanding of the native structure and interactions. While this has proven successful, the rational design of protein function is neither scalable nor generalizable, and manually optimizing proteins is difficult when the mechanistic basis of a protein’s activity are incomplete. To solve this challenged our lab has combined high-throughput laboratory experimentation with deep probabilistic modeling to enable the rapid optimization of protein function without requiring multiple experimental cycles. To date, we have applied out approach to a molecular chaperone, protease, RNA binding protein, and fluorescent protein. We are also engaged in multiple collaborations applying the approach to additional biomedically relevant targets.
Combinatorial protein engineering
Our lab previously developed a series of potent second generation Cas9-based transcriptional modulators. To date, these reagents have been requested by over 1,000 labs across the globe. In ongoing work, we have developed a novel method of screening combinatorial libraries of thousands of Cas9 variants to identify a third generation of Cas9 activators and repressors. Newly established effectors will be used to perform CRISPR screens to identify regulators of neuroinflammation and tumor cell growth. We are also adapting our regulators for in vivo applications within animal models of disease.
Synthetic gene drives to bias inheritance and aid in the dissection of fundamental biological processing
Gene drives are selfish genetic elements that "cheat" Mendelian inheritance. We were among the first to generate synthetic gene drives using Cas9. Our team is using this tool to perform rapid combinatorial knockouts within the clinically relevant fungal pathogen Candida albicans to understand the cellular mechanisms regulating pathogenicity.
DNA barcoded libraries for understanding regulators of neurodegeneration
By employing DNA barcodes, we are able to track the growth of dozens of cellular models at a time. We use these pools of barcoded cells to perform multiplexed screens aimed at understanding the genes involved in tolerance to misfolded proteins and investigating drugs that rescue various cell-based models of neurodegenerative disease. By using DNA-barcoding we are able to improve screening throughput by over an order of magnitude compared to conventional approaches, enabling us to obtain much richer datasets.
In vivo CRISPR screens
While CRISPR has revolutionized how we interrogate biological systems, to date much of the work comes from studying cells in isolation, outside of their physiologic environment. We have assembled an interdisciplinary team of scientist specializing in technology development, high-throughput screening, and neurobiology to perform in vivo CRISPR screens within animal models of neurodegeneration. By identifying genes which influence neuronal survival within neurodegenerative disease states, we aim to better understand the mechanisms underlying the profound neuronal loss seen within patients, along with identifying novel points for therapeutic intervention.
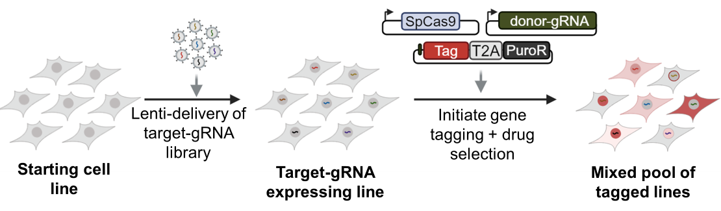
High-throughput genetic insertions
Our team has developed a strategy, High-throughput insertion of tags across the genome (HITAG), to perform precise genetic insertions into the mammalian genome at library scales. Using our HITAG approach, we are collaborating with multiple teams to characterize families of proteins involved in tumor survival, transcriptional regulation, and splicing. We are also further adapting the approach for performing other genetic modifications and for use in vivo to enable us to study protein function within native contexts.
Building technologies to speed the rate of antiviral drug discovery
Expanding upon our lab’s expertise in assay development, we have created a series of technologies to help increase the pace of antiviral drug discovery. These innovations have ranged from devising a method to screen for viral protease inhibitors in mammalian cells without requiring the use of live virus, to generating a novel multiplex drug screening approach that lets us search for inhibitors against dozen of viral proteins at once. Our long-term goal is to leverage these technologies to not only address the ongoing COVID-19 pandemic but also to help fill the numerous gaps within our existing antiviral armament.
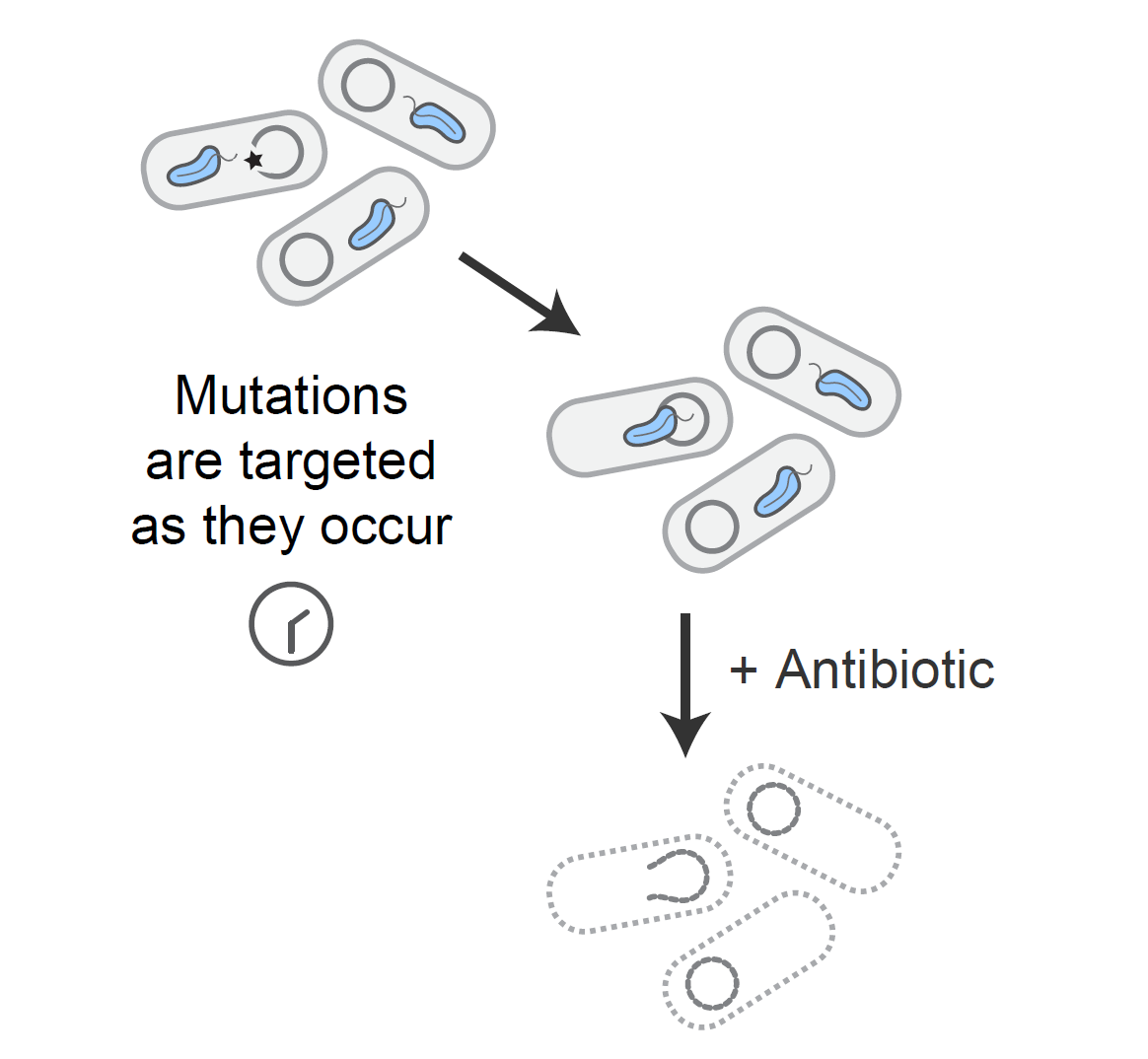
Tuned guide RNAs to control evolution
We developed a system to endow Cas9 with single nucleotide specificity through the development of tuned guide RNAs. With this method in hand, we developed a "genomic surveillance system" which prevents cells from gaining undesired mutations. We are exploring uses of this technology to control evolutionary trajectories to enrich for unconventional solutions to a given selection pressure.